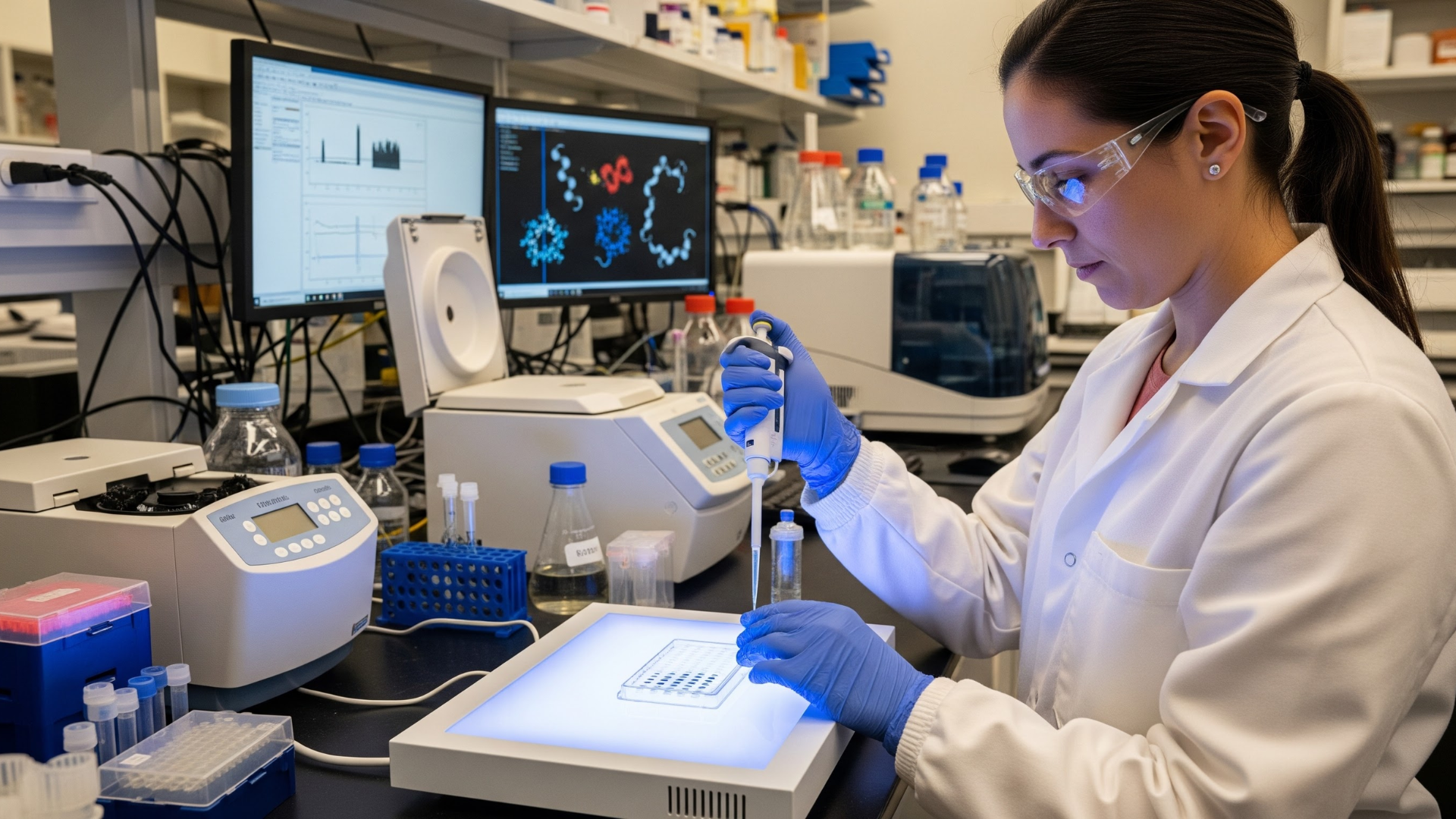
In many protein purification workflows, removing an affinity tag is supposed to be a routine final step. Yet for researchers working with the Tmprss15 recombinant protein, this step can quickly become unpredictable. The setup looks correct, the recognition sequence is present, and the incubation conditions follow standard recommendations. Still, the tag remains stubbornly attached.
This issue is more common than it appears. It often surfaces late in the workflow, after expression and purification have already gone well. That makes it particularly frustrating, especially when the downstream application depends on a clean, tag-free protein.
In most cases, the problem lies in how the substrate protein behaves under real conditions.
What factors cause TMPRSS15 cleavage to fail?
There are several factors that can affect how well the Tmprss15 recombinant protein performs. Let’s take a look at what might be going wrong with this thing.
- The Cleavage Site Is Not Truly Accessible
You may assume the DDDDK sequence is always easy for the Tmprss15 recombinant protein to recognize, but in real experiments, accessibility is often the main limitation. Proteins do not remain as open linear chains in solution. They fold into compact 3D structures, and this folding can bury the cleavage site deep inside the protein core.
So you put the tag in the place, but the enzyme still does not work. This happens a lot when you are working with proteins that are all folded up or have parts. It also happens when the tag is too close to a part of the protein that has a shape. The enzyme just cannot get to the tag in these situations. This is a problem with proteins that are tightly folded or have domains.
- Protein Folding Works Against The Reaction
When you work with the Tmprss15 recombinant protein, you may notice that even correctly designed constructs still resist cleavage. One major reason is protein folding stability. Folding does not just hide the cleavage site; it can lock the protein into a rigid shape that resists enzymatic access. Some proteins remain highly stable even under mild denaturing conditions, making them difficult substrates. Although native conditions are generally preferred to preserve enzyme activity, they may keep the substrate too rigid for efficient cleavage.
- Reaction Conditions Are Not As Optimal As They Seem
Even when protocols look correct on paper, small variations in conditions can strongly affect how the Tmprss15 recombinant protein performs. You should pay close attention to factors like pH, salt concentration, and calcium levels. Calcium plays a role because it helps keep the enzyme structure stable. If calcium levels are low, it can really cut down on the enzyme activity. A slight change in pH from the range can also make the enzyme less efficient at cutting.
The makeup of the buffers is also more important than you might think. High salt levels can help keep the protein folded correctly and make it harder for the enzyme to access its site. On the other hand, very low salt levels can make the protein unstable or cause it to clump together. Calcium and pH levels are crucial for enzyme activity.
The right buffer composition, including calcium and salt levels, helps the enzyme work properly.
- Enzyme Quality And Handling Matter More Than Expected
When working with the Tmprss15 recombinant protein, you may overlook enzyme handling as a cause of failure, but it plays a major role. Even if your enzyme looks fine, repeated freeze-thaw cycles can make a difference. This often leads to incomplete cleavage that seems like a substrate issue. You so what can you do to avoid loss of enzyme activity?
- Store the enzyme in small aliquots and keep conditions stable, preferably with glycerol.
- Check its activity on a control substrate before important experiments.
- The Enzyme-To-Substrate Ratio Is Not Always The Issue
When cleavage fails, you might instinctively increase the amount of Tmprss15 recombinant protein, assuming the enzyme concentration is too low. However, this is not always the real problem. The issue is that the cleavage site is hidden inside a tightly folded protein or blocked by its structure.
When the cleavage site is open, normal levels of the enzyme usually work fine.
Most of the time, making the construct design or making the linker more flexible fixes the problem, and that is more effective than just using more enzyme.
- Complex Samples Introduce Hidden Challenges
If you are working with partially purified proteins or cell lysates, Tmprss15 recombinant protein activity can be affected by several hidden factors. Protease inhibitors present in the sample may block enzyme function without obvious signs. In addition, other proteins in the mixture can interfere by competing for binding or altering the reaction environment.
The sample matrix has an impact on how well the reaction works. Sometimes the sample matrix can cause problems with the reaction, even when everything else seems to be fine. You might not think to look at the sample matrix. It is really important in experiments. To get results, you should think about cleaning up the sample before you start the reaction.
This can be something like getting rid of things that can stop the reaction or changing the buffer solution that the sample is in. The sample matrix can reduce how well the reaction works, so it is a good idea to consider the sample matrix when you are setting up your experiment.